J Clin Invest | Jin-Xin Bei’s Team Uncovers a Super-Enhancer–Driven Mechanism Underlying Malignant Progression of Cervical Cancer
Cervical cancer (CC) remains a major global public health concern, ranking as the fourth most prevalent cancerand the fourth leading cause of cancer-related mortality among womenworldwide. The pathogenesis of CC is a complex, multistep process driven by persistent infection with high-risk human papillomavirus (HPV) subtypes, in conjunction with host genetic susceptibility and environmental factors.Although early-stage CC can often be effectively managed through radical surgery or chemoradiotherapy, the majority of patients diagnosed at advanced stages, particularly those with recurrent or metastatic disease, exhibit poor clinical outcomes, with a 5-year survival rate as low as 16.5%. Accordingly, elucidating the molecular mechanisms underlying cervical carcinogenesis is critical for the development of more effective diagnostic and therapeutic strategies.
Super-enhancers (SEs) are large clusters of transcriptional enhancers distinguished by exceptionally high levels of enhancer-associated histone modifications and dense occupancy by transcriptional co-activators and mediator complexes. Accumulating evidence indicates that SEs play pivotal roles in driving oncogene expression across multiple cancer types, including CC. However, current understanding of SE-mediated regulation in CC is largely based on studies using established commercial cell lines, which may not accurately reflect the epigenetic architecture of primary tumors due to technical constraints and limited availability of clinical specimens.Importantly, substantial disparities in epigenetic landscapes have been observed between primary tumors and cell lines, underscoring the necessity for systematic, genome-wide characterization of SE landscapes in CC tumors and in-depth elucidation of the molecular mechanisms underlying their regulatory functions.
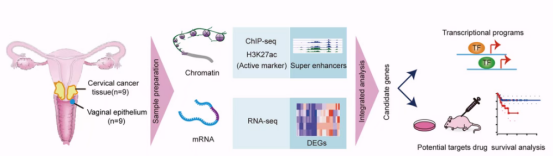
Recently, Prof. Jin-Xin Bei and colleagues at Sun Yat-sen University Cancer Center and Institute of Advanced Studies Hong Kong, Sun Yat-sen University, reported a study entitled “Super-Enhancer-Driven EFNA1 Fuels Tumor Progression in Cervical Cancer via the FOSL2–Src/AKT/STAT3 Axis” in the Journal of Clinical Investigation. By integrative analysis of chromatin immunoprecipitation sequencing (ChIP-seq) and RNA sequencing (RNA-seq) data derived from CC specimens and matched normal tissues, the research team constructed a CC–specific SE landscape. Functional and mechanistic studies using in vitro and in vivo models demonstrated that a tumor-specific EFNA1-associated SE drives aberrant EFNA1 overexpression, thereby promoting CC progression. Furthermore, the study evaluated the therapeutic potential of targeting EFNA1-mediated downstream signaling pathwaysin CC treatment.

In this study, the researchers performed an integrative multi-omics analysis incorporating H3K27acChIP-seq and RNA-seq data from nine paired CC tissues and matched normal controls, thereby, for the first time, establishing a comprehensive tissue-level SE landscape specific to CC. This analysis identified a subset of SE-associated genes that were selectively activated in tumor tissues (Figure 1). Subsequent survival analyses using The Cancer Genome Atlas (TCGA) cohort revealed EFNA1 as a key SE-driven gene, with elevated EFNA1 expression significantly associated with unfavorable patient prognosis (hazard ratio = 2.0, P = 0.0047), underscoring its critical role in CC progression.
Using CRISPR/Cas9 editingsystem, the researchersspecifically deleted the core enhancer elements (E1 and E2) within the EFNA1-SE region. Notably, loss of these elements resulted in substantial suppression of EFNA1 transcription, with mRNA levels reduced by 51% and 63%, respectively. Dual-luciferase reporter assays confirmed that E1 or E2 markedly enhanced EFNA1 promoter activity, an effect that was abrogated by the BRD4 inhibitor JQ-1. Mechanistically, the EFNA1-SE is highly enriched for H3K27ac and recruits the transcription factor FOSL2, which directly binds to the SE region and cooperates with H3K27ac to drive EFNA1 transcription. Functional studies demonstrated that disruption of the EFNA1-SE or EFNA1 silencing significantly impaired cervical cancer cell proliferation, migration, invasion, and tumorigenic potential, whereas EFNA1 overexpression enhanced these malignant phenotypes. Collectively, these findings establish the SE-driven EFNA1 as akey determinant of CC aggressiveness.
To further delineate the oncogenic mechanisms mediated by EFNA1, the researchers integrated RNA-seq, protein microarray profiling, and co-immunoprecipitation (Co-IP) assays. These analyses revealed that EFNA1 engages in cis-interaction with the FNIII domain of the EphA2 receptor, leading toreduced EphA2 phosphorylation and increased proteasomal degradation. This regulatory interaction subsequently results in aberrant activation of the downstream Src/AKT/STAT3 signaling cascade, thereby driving CC malignant progression. Importantly, pharmacological inhibition of key molecules within this pathway, including Src inhibition by Saracatinib, Akt inhibition by MK2206, and STAT3 inhibition by Static, effectively abrogated the pro-tumorigenic effects induced by EFNA1 overexpression.
In summary, this study delineates a CCtumor–specific SE regulatory landscape and identifies EFNA1 as a previously unrecognized SE–driven oncogene. Furthermore, it elucidates the molecular mechanism underlying activation of the EFNA1–EphA2–Src/AKT/STAT3 signaling axis and demonstrates its critical role in drivingCC progression (Figure 2). These findings advance our understanding of epigenetic regulation in CC and provide a strong conceptual framework for the development of therapeutic strategies targeting SEs and their associated downstream signaling pathways.

Dr. Shu-Qiang Liu, Dr. Xi-Xi Cheng, and Dr. Shuai He from Sun Yat-sen University Cancer Center contributed equally to this work as co–first authors. Prof. Jin-Xin Bei, Dr. Chun-Ling Luo, and Prof. Chun-Yan Lan from Sun Yat-sen University Cancer Center are the corresponding authors.
Original article link:https://www.jci.org/articles/view/177599
Prof. Jin-Xin Bei’s research group is dedicated to precision and translational medicine, with a primary focus on systemic immunity and tumor heterogeneity. By integrating multi-omics datasets, including genomic, epigenetic, and transcriptomic data at both bulk and single-cell resolutions, the group systematically dissects cellular interactions and regulatory networks within the tumor microenvironment that drive tumor initiation and progression. Through these efforts, the team seeks to identify reliable biomarkers for early and accurate cancer diagnosis and to discover actionable therapeutic targets for precise intervention, ultimately advancing personalized strategies for cancer prevention and treatment.
To date, the group has published over 130 SCI-indexed articles in leading international journals, including Cell, The Lancet Oncology, Nature Genetics, Journal of Clinical Investigation, and Nature Communications. The group actively recruits outstanding scientists at multiple career stages and welcomes highly motivated postdoctoral fellows and doctoral students with strong research interests in related fields to join the team.

