Cell: A Multi-Organ Immune Cell Atlas Reveals New Mechanisms Underlying Systemic Immune Establishment and Tolerance
On October 28, 2025, a collaborative team led by ProfessorJin-XinBei from Sun Yat-sen UniversityInstitute of Advanced Studies published an article entitled “Systemic immune activity occurs during human immune system maturation” in Cell.Based on 321 samples from 23 fetal and adult tissues, this study analyzed more than 2.8 million immune cells using single-cell transcriptomics and TCR sequencing, providing the first systematic and dynamic atlas of human immune system development during the second trimester. The authors discovered widespread and orderly systemic activation of fetal T cells, identified two previously unrecognized immune tolerance mechanisms, and revealed that hematopoietic activity is broadly distributed across multiple fetal organs. These findings illuminate the transition of the immune system from local to systemic maturation, fundamentally revising the long-held concept of “fetal immune quiescence” and offering theoretical insights relevant to organ transplantation and cancer therapy optimization.
The establishment of systemic immunity is closely intertwined with human organ development. Traditionally, the fetal immune system was thought to remain largely quiescent in utero. However, emerging evidence indicates that fetal immune cells—particularly T cells—already possess functional responsiveness, challenging this paradigm. Meanwhile, fetal development must proceed under strict regulation of maternal-fetal immune tolerance to avoid detrimental inflammation. Although seemingly paradoxical, this phenomenon reflects the remarkable complexity of cross-organ coordination and systemic regulation within the fetal immune system. These observations underscore the importance of studying immune development from a system-level perspective, focusing on key dynamic events and tolerance mechanisms that support healthy embryogenesis.
To compare fetal and adult immune systems, the research team focused on the second trimester (11~26 weeks post-conception), a critical developmental window. They collected samples from 15 aborted human fetal donors and 4 adult donors, encompassing 23 distinct organs. By enriching immune cells and integrating single-cell transcriptomic and TCR sequencing, the authors generated the first multi-organ, within-individual systemic immune atlas spanning both fetal and adult stages. Analytical results revealed the presence of activated and memory-like T cells across multiple fetal organs, accompanied by TCR clonal expansion. Notably, the proportion of activated/effector T cells increased progressively with gestational age, indicating that fetal immune responses are not confined to individual organs but instead involve coordinated systemic activation. Further TCR analyses demonstrated extensive clonal sharing among adult organs, particularly among tissue-resident memory T cells (TRM) in digestive system-associated tissues. These findings challenge conventional views by suggesting that TRM cells participate not only in local barrier defense but also in the broader systemic immune network, jointly regulating immunity across circulating and resident compartments.
Despite fetal T cells exhibited memory features and activated phenotypes, they showed markedly lower activation levels than adult T cells, indicating the existence of regulatory mechanisms that restrain excessive activation and maintain immune homeostasis during development. The study identified abundant immature neutrophils across multiple fetal organs that highly express the immunosuppressive molecule ARG1. ARG1 expression was consistently high before birth but declined to minimal levels in adulthood. Co-culture experiments demonstrated that both fetal and adult T cells were strongly suppressed by fetal neutrophils via ARG1, whereas adult neutrophils exerted little effect. These results establish immature fetal neutrophils as the primary in vivo source of ARG1, functioning as a “buffer layer” to prevent excessive T-cell activation and thereby maintain immune tolerance during development.
Building on these findings, the authors further investigated how intercellular signaling regulates T-cell activation. Interaction network analyses showed that nearly all fetal cell types could signal to quiescent fetal T cells through the PTGES3 (PGE2)-PTGER4 ligand-receptor pathway, with markedly stronger activity in fetuses than in adults. Co-culture experiments confirmed that blocking this pathway effectively relieved suppression of T-cell activation. Together, these data demonstrate that the fetal microenvironment systemically delivers inhibitory signals to T cells via the PTGES3 (PGE2)-PTGER4 axis, representing a key mechanism of immune tolerance that operates in concert with ARG1⁺ neutrophils to shape a global tolerance network.
Although the liver is widely regarded as the primary hematopoietic organ during fetal development, this study detected hematopoietic stem cells (HSCs) in multiple organs, including the heart, skin, kidney, spleen, peripheral blood, and intestine. The proportion of HSCs among immune cells declined with advancing gestational age. Immunofluorescence analyses confirmed that HSCs were predominantly located within tissue parenchyma rather than vascular lumens, excluding the possibility that they merely reflected residual circulating blood. Functional assays showed that both liver-derived HSCs and those from other organs-exemplified by the heart-possessed multilineage differentiation potential. Liver HSCs preferentially differentiated into monocytes/macrophages and neutrophils, whereas cardiac HSCs favored erythroid and mast-cell lineages. These results reveal a cross-organ hematopoietic network in the fetus, providing multiple spatial sources for immune cell supply and supporting systemic immune maturation.
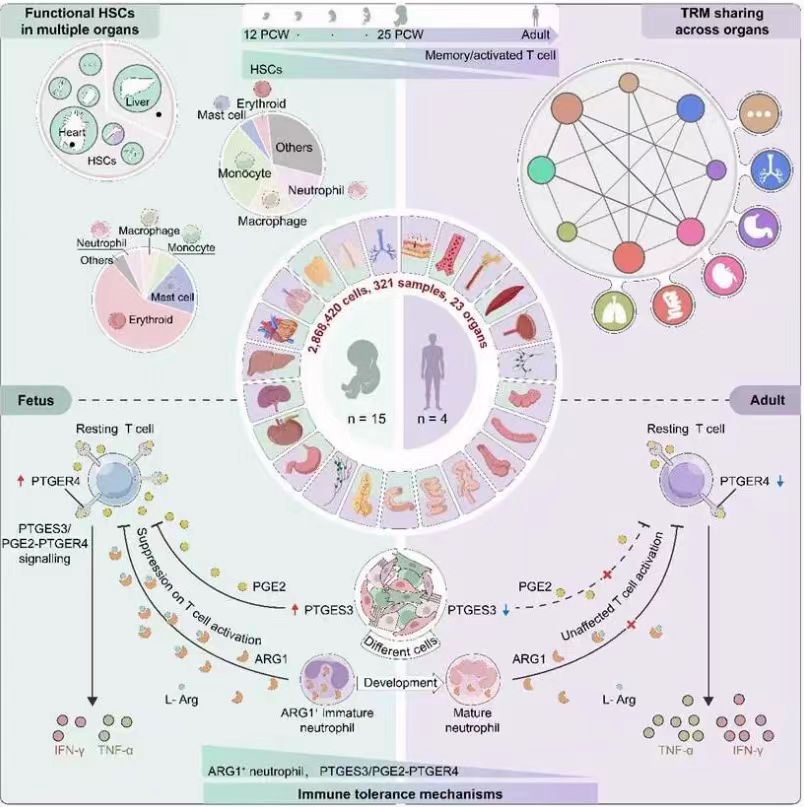
In summary, this study presents the first dynamic atlas of human immune system development, uncovering systemic activation and clonal expansion of fetal T cells, coordinated immune tolerance mediated by ARG1 and the PTGES3-PTGER4 pathway, and multi-organ HSC-driven spatial hematopoiesis. By framing immune development through the lens of “systemic immunity,” these findings advance our understanding of immune establishment and tolerance and open new avenues for preventing and treating complex diseases such as cancer.
Professors Xiao-shun He, Jin-xin Bei, Zhi-yong Guo, and Zi-lian Wang from Sun Yat-sen University served as corresponding authors. Associate Professors Shuai He and Chun-ling Luo (Sun Yat-sen University Cancer Center), Dr. Luo Tao (The First Affiliated Hospital of Sun Yat-sen University), and Professor Hai-tan Chen (Deputy Director, Department of Obstetrics) were co-first authors. Professors Xiao-yi Wang (The Third Affiliated Hospital of Guangzhou Medical University), Qi-tao Huang (Foshan First People’s Hospital), Yu-jie Gan (Boai Hospital of Zhongshan, Maternal and Child Health Hospital), and Associate Professor Hai-hua Zhang (Baiyun District Maternal and Child Health Hospital, Guangzhou) provided critical support. This work was jointly conducted by Sun Yat-sen University and the Hong Kong Institute of Advanced Studies at Sun Yat-sen University, with technical support from the High-Throughput Sequencing Platform of Sun Yat-sen University Cancer Center.
Original article:https://www.cell.com/cell/fulltext/S0092-8674(25)01132-8
Prof. Jin-Xin Bei’s research group is dedicated to precision and translational medicine, with a primary focus on systemic immunity and tumor heterogeneity. By integrating multi-omics datasets, including genomic, epigenetic, and transcriptomic data at both bulk and single-cell resolutions, the group systematically dissects cellular interactions and regulatory networks within the tumor microenvironment that drive tumor initiation and progression. Through these efforts, the team seeks to identify reliable biomarkers for early and accurate cancer diagnosis and to discover actionable therapeutic targets for precise intervention, ultimately advancing personalized strategies for cancer prevention and treatment.
To date, the group has published over 130 SCI-indexed articles in leading international journals, including Cell, The Lancet Oncology, Nature Genetics, Journal of Clinical Investigation, and Nature Communications. The group actively recruits outstanding scientists at multiple career stages and welcomes highly motivated postdoctoral fellows and doctoral students with strong research interests in related fields to join the team.

